Code
# Load packages and data
library(shiny)
library(tidyverse)
library(tidymodels)
library(kknn)
library(ISLR)
data(College)
college_demo <- College %>%
mutate(school = rownames(College)) %>%
filter(Grad.Rate <= 100)shiny and kknn packages if you haven’t already
When:
Topics:
Format:
set_mode) or argument (eg "regression")step_dummy() function do?)step_dummy() function?)
HW3 (due Friday, Feb 27):
Group Assignment 1 (due Friday, Mar 6):
Other resources (use, on your own, however you find useful):
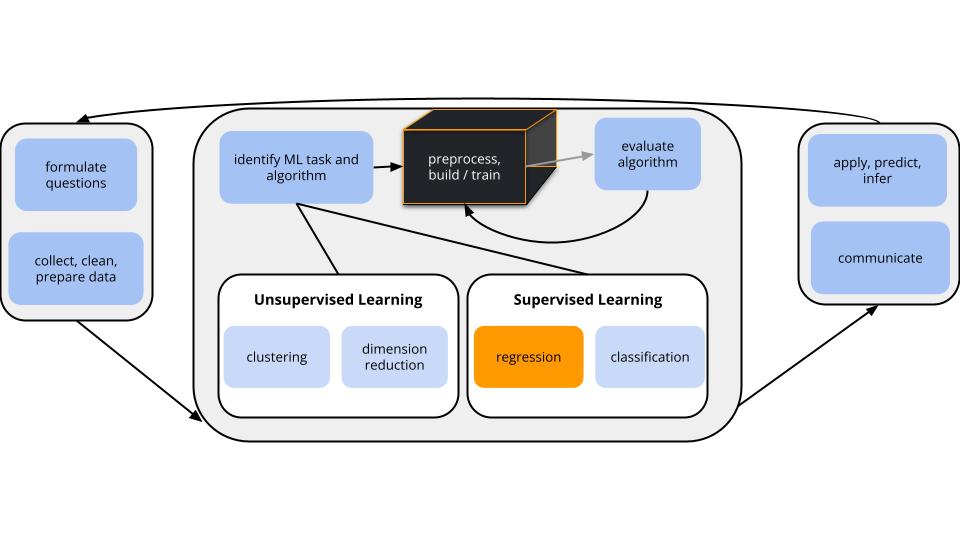
world = supervised learning
We want to model some output variable \(y\) using a set of potential predictors (\(x_1, x_2, ..., x_p\)).
task = regression
\(y\) is quantitative
(nonparametric) algorithm = K Nearest Neighbors (KNN)
Our usual parametric models (eg: linear regression) are too rigid to represent the relationship between \(y\) and our predictors \(x\). Thus we need more flexible nonparametric models.
Goal
Build a flexible regression model of a quantitative outcome \(y\) by a set of predictors \(x\),
\[y = f(x) + \varepsilon\]
. . .
Idea
Predict \(y\) using the data on “neighboring” observations. Since the neighbors have similar \(x\) values, they likely have similar \(y\) values.
. . .
Algorithm
For tuning parameter K, take the following steps to estimate \(f(x)\) at each set of possible predictor values \(x\):
. . .
Output
KNN does not produce a nice formula for \(\hat{f}(x)\), but rather a set of rules for how to calculate \(\hat{f}(x)\).
. . .
In pictures (from ISLR)
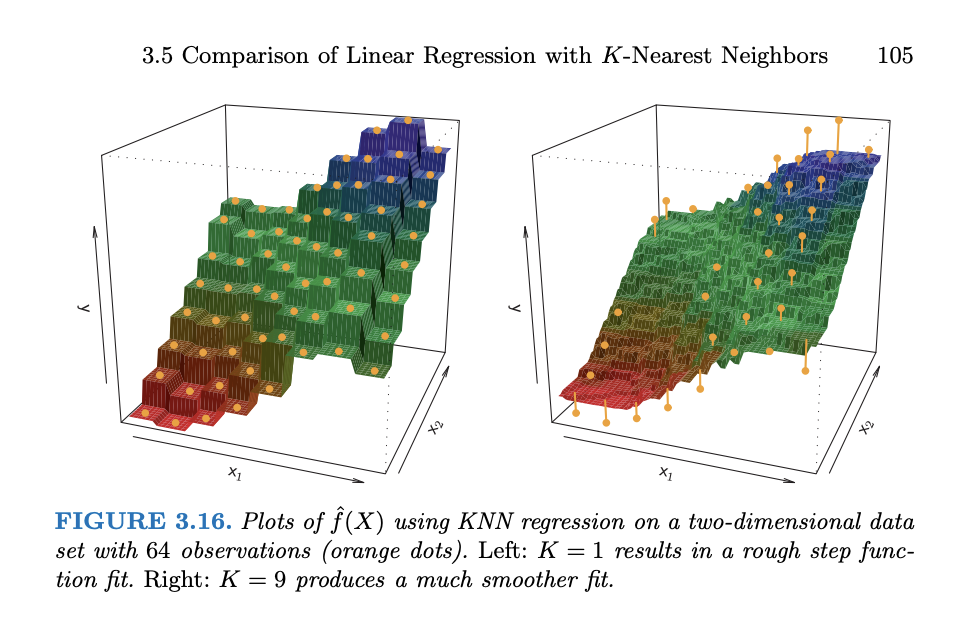
Discuss the following questions with your group.
Let’s review the KNN algorithm using a shiny app. Run the code below and ignore the syntax!!
Click “Go!” one time only to collect a set of sample data.
Check out the KNN with K = 1.
Now try the KNN with K = 25.
Set K = 100 where 100 is the number of data points. Is this what you expected?
# Load packages and data
library(shiny)
library(tidyverse)
library(tidymodels)
library(kknn)
library(ISLR)
data(College)
college_demo <- College %>%
mutate(school = rownames(College)) %>%
filter(Grad.Rate <= 100)# Define a KNN plotting function
plot_knn <- function(k, plot_data){
expend_seq <- sort(c(plot_data$Expend, seq(3000, 57000, length = 5000)))
#knn_mod <- knn.reg(train = plot_data$Expend, test = data.frame(expend_seq), y = plot_data$Grad.Rate, k = k)
knn_results <- nearest_neighbor() %>%
set_mode("regression") %>%
set_engine(engine = "kknn") %>%
set_args(neighbors = k) %>%
fit(Grad.Rate ~ Expend, data = plot_data) %>%
augment(new_data = data.frame(Expend = expend_seq)) %>%
rename(expend_seq = Expend, pred_2 = .pred)
ggplot(plot_data, aes(x = Expend, y = Grad.Rate)) +
geom_point() +
geom_line(data = knn_results, aes(x = expend_seq, y = pred_2), color = "red") +
labs(title = paste("K = ", k), y = "Graduation Rate", x = "Per student expenditure ($)") +
lims(y = c(0,100))
}
# BUILD THE SERVER
# These are instructions for building the app - what plot to make, what quantities to calculate, etc
server_KNN <- function(input, output) {
new_data <- eventReactive(input$do, {
sample_n(college_demo, size = 100)
})
output$knnpic <- renderPlot({
plot_knn(k = input$kTune, plot_data = new_data())
})
}
# BUILD THE USER INTERFACE (UI)
# The UI controls the layout, appearance, and widgets (eg: slide bars).
ui_KNN <- fluidPage(
sidebarLayout(
sidebarPanel(
h4("Sample 100 schools:"),
actionButton("do", "Go!"),
h4("Tune the KNN algorithm:"),
sliderInput("kTune", "K", min = 1, max = 100, value = 1)
),
mainPanel(
h4("KNN Plot:"),
plotOutput("knnpic")
)
)
)
# RUN THE SHINY APP!
shinyApp(ui = ui_KNN, server = server_KNN)
The tidymodels KNN algorithm predicts \(y\) using weighted averages. The idea is to give more weight or influence to closer neighbors, and less weight to “far away” neighbors.
More details in the kknn package documentation.
Optional math:
Let (\(y_1, y_2, ..., y_K\)) be the \(y\) outcomes of the K neighbors and (\(w_1, w_2, ..., w_K\)) denote the corresponding weights. These weights are defined by a “kernel function” which ensures that: (1) the \(w_i\) add up to 1; and (2) the closer the neighbor \(i\), the greater its \(w_i\). Then the neighborhood prediction of \(y\) is:
\[\sum_{i=1}^K w_i y_i\] The kernel wikipedia page shows common kernel functions.
The default “optimal” kernel used in kknn was defined in a 2012 Annals of Statistics paper.
What would happen if we had gotten a different sample of data?!?
In general…
Why is “high bias” bad?
Why is “high variability” bad?
What is meant by the bias-variance tradeoff?
Be kind to yourself (and each other). You will make mistakes! That’s part of the learning process.
Stay engaged. Studies show that when you’re working on an assignment for another class, continuously on your message app, playing games, etc. it impacts both your learning and the learning of those around you.
Using the College dataset from the ISLR package, we’ll explore the KNN model of college graduation rates (Grad.Rate) by:
Expend)Enroll)Private# Load packages
library(tidymodels)
library(tidyverse)
library(ISLR)
# Load data
data(College)
# Wrangle the data
college_sub <- College %>%
mutate(school = rownames(College)) %>%
arrange(factor(school, levels = c("Macalester College", "Luther College", "University of Minnesota Twin Cities"))) %>%
filter(Grad.Rate <= 100) %>%
filter((Grad.Rate > 50 | Expend < 40000)) %>%
select(Grad.Rate, Expend, Enroll, Private)Check out a codebook from the console:
?College
Goals
The KNN model for Grad.Rate will hinge upon the neighborhoods defined by the 3 Expend, Enroll, and Private predictors. And these neighborhoods hinge upon how we pre-process our predictors.
We’ll explore these ideas below using the results of the following chunk. Run this, but DON’T spend time examining the code!
recipe_fun <- function(recipe){
recipe <- recipe %>%
prep() %>%
bake(new_data = college_sub) %>%
head(3) %>%
select(-Grad.Rate) %>%
as.data.frame()
row.names(recipe) <- c("Mac","Luther", "UMN")
return(recipe)
}
# Recipe 1: create dummies, but don't standardize
recipe_1 <- recipe(Grad.Rate ~ Expend + Enroll + Private, data = college_sub) %>%
step_nzv(all_predictors()) %>%
step_dummy(all_nominal_predictors())
recipe_1_data <- recipe_fun(recipe_1)
# Recipe 2: standardize, then create dummies
recipe_2 <- recipe(Grad.Rate ~ Expend + Enroll + Private, data = college_sub) %>%
step_nzv(all_predictors()) %>%
step_normalize(all_numeric_predictors()) %>%
step_dummy(all_nominal_predictors())
recipe_2_data <- recipe_fun(recipe_2)
# Recipe 3: create dummies, then standardize
recipe_3 <- recipe(Grad.Rate ~ Expend + Enroll + Private, data = college_sub) %>%
step_nzv(all_predictors()) %>%
step_dummy(all_nominal_predictors()) %>%
step_normalize(all_numeric_predictors())
recipe_3_data <- recipe_fun(recipe_3)
Check out the feature space of our 3 predictors and take note of which school is the closer neighbor of Mac: UMN or Luther.
ggplot(college_sub, aes(x = Expend, y = Enroll, color = Private)) +
geom_point(alpha = 0.5) +
geom_text(data = head(college_sub, 3), aes(x = Expend, y = Enroll, label = c("Mac","Luther", "UMN")), color = "black")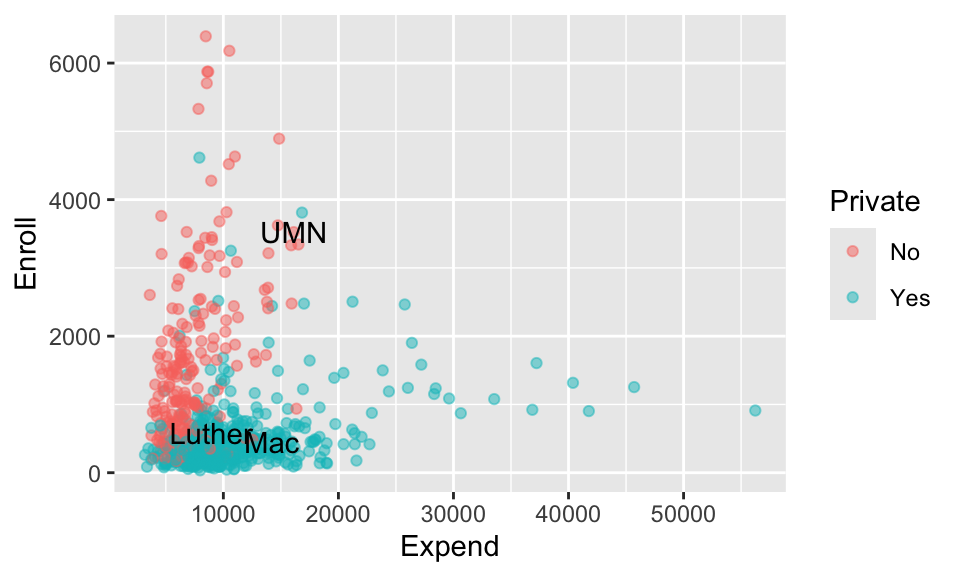
Of course, KNN relies upon mathematical metrics (Euclidean distance), not visuals, to define neighborhoods. And these neighborhoods depend upon how we pre-process our predictors. Consider the pre-processing recipe_1 which uses step_dummy() but not step_normalize():
recipe_1_data Expend Enroll Private_Yes
Mac 14213 452 1
Luther 8949 587 1
UMN 16122 3524 0sqrt((14213 - ___)^2 + (452 - ___)^2 + (1 - ___)^2)dist().dist(recipe_1_data)
recipe_2 first uses step_normalize() and then step_dummy() to pre-process the predictors:recipe_2_data Expend Enroll Private_Yes
Mac 0.9025248 -0.3536904 1
Luther -0.1318021 -0.2085505 1
UMN 1.2776255 2.9490477 0Calculate the distance between each pair of schools using these pre-processed data:
dist(recipe_2_data)By this metric, is Mac closer to Luther or UMN? So is this a reasonable metric?
recipe_2 first uses step_normalize() and then step_dummy() to pre-process the predictors, recipe_3 first uses step_dummy() and then step_normalize():recipe_3_data Expend Enroll Private_Yes
Mac 0.9025248 -0.3536904 0.6132441
Luther -0.1318021 -0.2085505 0.6132441
UMN 1.2776255 2.9490477 -1.6285680How do the pre-processed data from recipe_3 compare those to recipe_2?
RECALL: The standardized dummy variables lose some contextual meaning. But, in general, negative values correspond to 0s (not that category), positive values correspond to 1s (in that category), and the further a value is from zero, the less common that category is.
Calculate the distance between each pair of schools using these pre-processed data. By this metric, is Mac closer to Luther or UMN?
dist(recipe_3_data)recipe_3 compare to those from recipe_2?dist(recipe_2_data)recipe_2, recipe_3 considered the fact that private schools are relatively more common in this dataset, making the public UMN a bit more unique. Why might this be advantageous when defining neighborhoods? Thus why will we typically first use step_dummy() before step_normalize()?college_sub %>%
count(Private)
With a grip on neighborhoods, let’s now build a KNN model for Grad.Rate.
For the purposes of this activity (focusing on concepts over R code), simply run each chunk and note what object it’s storing.
You will later be asked to come back and comment on the code.
STEP 1: Specifying the KNN model
knn_spec <- nearest_neighbor() %>%
set_mode("regression") %>%
set_engine(engine = "kknn") %>%
set_args(neighbors = tune())STEP 2: Variable recipe (with pre-processing)
Note that we use step_dummy() before step_normalize().
variable_recipe <- recipe(Grad.Rate ~ ., data = college_sub) %>%
step_nzv(all_predictors()) %>%
step_dummy(all_nominal_predictors()) %>%
step_normalize(all_numeric_predictors())STEP 3: workflow specification (model + recipe)
knn_workflow <- workflow() %>%
add_model(knn_spec) %>%
add_recipe(variable_recipe)STEP 4: estimate multiple KNN models
This code builds 50 KNN models of Grad.Rate, using 50 possible values of K ranging from 1 to 200 (roughly 25% of the sample size of 775).
It then evaluates these models with respect to their 10-fold CV MAE.
set.seed(253)
knn_models <- knn_workflow %>%
tune_grid(
grid = grid_regular(neighbors(range = c(1, 200)), levels = 50),
resamples = vfold_cv(college_sub, v = 10),
metrics = metric_set(mae)
)
knn_models %>%
autoplot()
best_k <- knn_models %>%
select_best(metric = "mae")
best_k# Plug in a number or best_k$neighbors
knn_models %>%
collect_metrics() %>%
filter(neighbors == ___)
Build your “final” KNN model using the optimal value you found for K above. NOTE: We only looked at roughly every 4th possible value of K (K = 1, 5, 9, etc). If we wanted to be very thorough, we could re-run our algorithm using each value of K close to our optimal K.
final_knn <- knn_workflow %>%
finalize_workflow(parameters = best_k) %>%
fit(data = college_sub)What does a tidy() summary of final_knn give you? Does this surprise you?
# DO NOT REMOVE eval = FALSE
final_knn %>%
tidy()
# Check out Mac & Luther
mac_luther <- college_sub %>%
head(2)
mac_luther Grad.Rate Expend Enroll Private
Macalester College 77 14213 452 Yes
Luther College 77 8949 587 Yes# Prediction
___ %>%
___(new_data = ___)
What assumptions did the KNN model make about the relationship of Grad.Rate with Expend, Enroll, and Private?
What did the KNN model tell us about the relationship of Grad.Rate with Expend, Enroll, and Private?
Reflecting upon a and b, name one pro of using a nonparametric algorithm like the KNN instead of a parametric algorithm like least squares or LASSO.
Similarly, name one con.
Consider another “con”. Just as with parametric models, we could add more and more predictors to our KNN model. However, the KNN algorithm is known to suffer from the curse of dimensionality. Why? (A quick Google search might help.)
ggplot(College, aes(x = Outstate, y = Room.Board)) +
geom_point()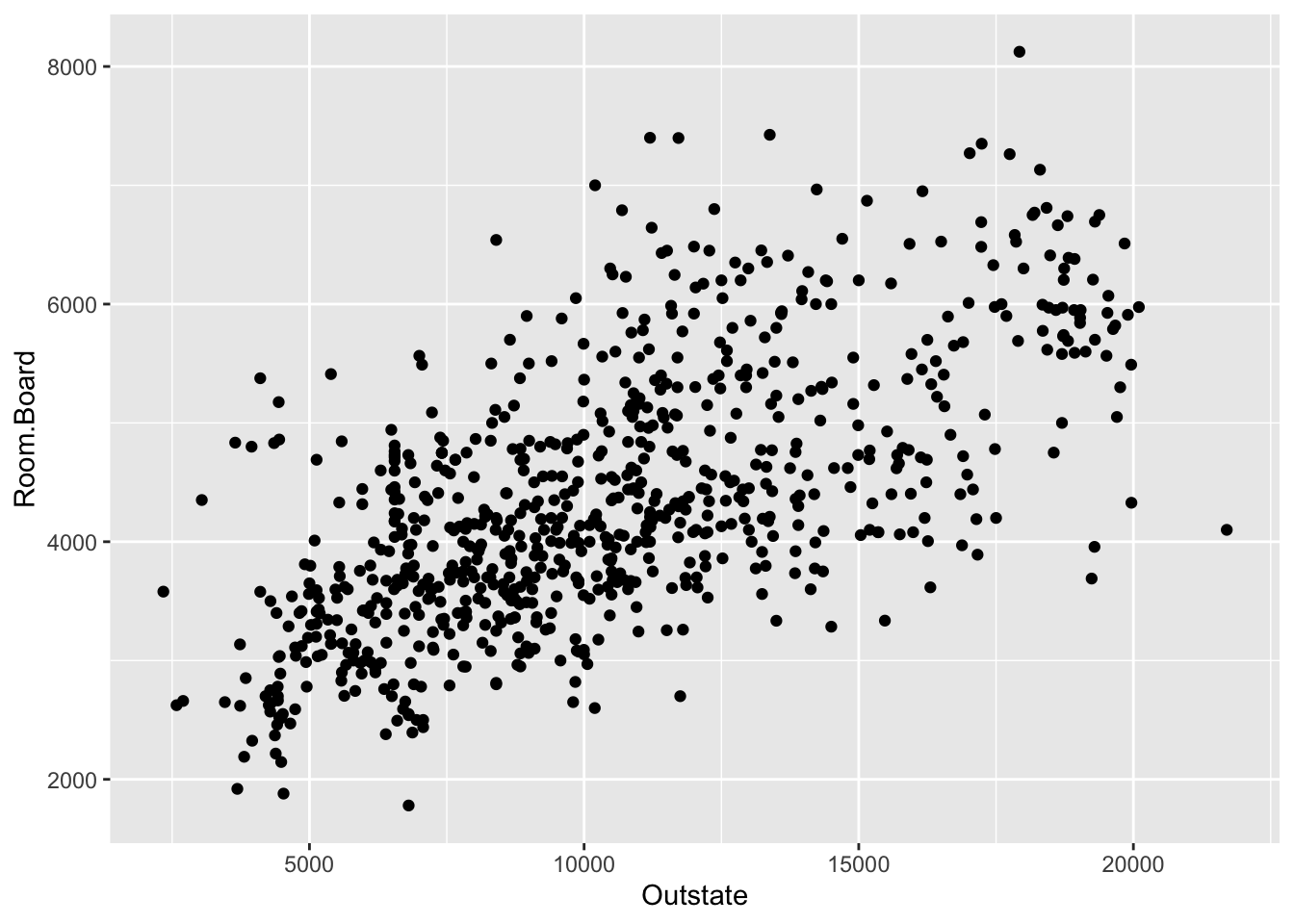
Revisit all code in Parts 2 and 3 of the exercises. Comment upon each chunk:
We previously used select_best() to choose the number of neighbors \(k\) that resulted in the lowest CV MAE.
Recall from our LASSO activity that we can also use select_by_one_std_err() to choose most simple model that produces a CV MAE that’s within 1 standard error of the best model (thus is not significantly worse). Review the code below:
# get "parsimonous" lambda
parsimonious_penalty <- lasso_models %>%
select_by_one_std_err(metric = "mae", desc(penalty))Let’s figure out how to adapt this to the KNN setting.
Enter ?select_by_one_std_err in the Console to pull up the help page for this function.
Scroll down to the Arguments section, and note what the ... argument does.
Use select_by_one_std_err() to store the parsimonious choice for \(k\) as parsimonious_k (or simple_k). (Our tuning parameter is called neighbors in tidymodels.)
Use knn_models %>% collect_metrics() %>% filter(neighbors == ___) to compare the CV MAE for the best \(k\) to the \(k\) for the simpler model.
Enroll for public vs private schools.Grad.Rate vs Enroll, Private, and Expend.
Suppose we want to build a model of response variable y using predictors x1 and x2 in our sample_data.
# Load packages
library(tidymodels)
library(kknn)
Build the model
# STEP 1: KNN model specification
knn_spec <- nearest_neighbor() %>%
set_mode("regression") %>%
set_engine(engine = "kknn") %>%
set_args(neighbors = tune())STEP 1 notes:
kknn, not lm, engine to build the KNN model.knn engine requires us to specify an argument (set_args):
neighbors = tune() indicates that we don’t (yet) know an appropriate value for the number of neighbors \(K\). We need to tune it.set_args().# STEP 2: variable recipe
# (You can add more pre-processing steps.)
variable_recipe <- recipe(y ~ x1 + x2, data = sample_data) %>%
step_nzv(all_predictors()) %>%
step_dummy(all_nominal_predictors()) %>%
step_normalize(all_numeric_predictors())# STEP 3: workflow specification (model + recipe)
knn_workflow <- workflow() %>%
add_recipe(variable_recipe) %>%
add_model(knn_spec)# STEP 4: Estimate multiple KNN models using a range of possible K values
# Calculate the CV MAE & R^2 for each
set.seed(___)
knn_models <- knn_workflow %>%
tune_grid(
grid = grid_regular(neighbors(range = c(___, ___)), levels = ___),
resamples = vfold_cv(sample_data, v = ___),
metrics = metric_set(mae, rsq)
)STEP 4 notes:
set.seed(___).tune_grid() instead of fit() since we have to build multiple KNN models, each using a different tuning parameter K.grid specifies the values of tuning parameter K that we want to try.
range specifies the lowest and highest numbers we want to try for K (e.g. range = c(1, 10). The lowest this can be is 1 and the highest this can be is the size of the smallest CV training set.levels is the number of K values to try in that range, thus how many KNN models to build.resamples and metrics indicate that we want to calculate a CV MAE for each KNN model.
Tuning K
# Calculate CV MAE for each KNN model
knn_models %>%
collect_metrics()
# Plot CV MAE (y-axis) for the KNN model from each K (x-axis)
autoplot(knn_models)
# Identify K which produced the lowest ("best") CV MAE
best_K <- select_best(knn_models, metric = "mae")
best_K
# Get the CV MAE for KNN when using best_K
knn_models %>%
collect_metrics() %>%
filter(neighbors == best_K$neighbors)
Finalizing the “best” KNN model
# parameters = final K value (best_K or whatever other value you might want)
final_knn_model <- knn_workflow %>%
finalize_workflow(parameters = ___) %>%
fit(data = sample_data)
Use the KNN to make predictions
# Put in a data.frame object with x1 and x2 values (at minimum)
final_knn_model %>%
predict(new_data = ___)
# a
sqrt((14213 - 8949)^2 + (452 - 587)^2 + (1 - 1)^2)[1] 5265.731# b
dist(recipe_1_data) Mac Luther
Luther 5265.731
UMN 3616.831 7750.993Luther. Yes.
recipe_2_data Expend Enroll Private_Yes
Mac 0.9025248 -0.3536904 1
Luther -0.1318021 -0.2085505 1
UMN 1.2776255 2.9490477 0dist(recipe_2_data) Mac Luther
Luther 1.044461
UMN 3.471135 3.599571recipe_3_data Expend Enroll Private_Yes
Mac 0.9025248 -0.3536904 0.6132441
Luther -0.1318021 -0.2085505 0.6132441
UMN 1.2776255 2.9490477 -1.6285680Private_Yes is now 0.6132441 or -1.6285680, not 1 or 0.
Luther
dist(recipe_3_data) Mac Luther
Luther 1.044461
UMN 4.009302 4.120999dist(recipe_2_data) Mac Luther
Luther 1.044461
UMN 3.471135 3.599571
No “answers” here. Just run the provided code and make note of what it’s doing.
STEP 1: Specifying the KNN model
knn_spec <- nearest_neighbor() %>%
set_mode("regression") %>%
set_engine(engine = "kknn") %>%
set_args(neighbors = tune())STEP 2: Variable recipe (with pre-processing)
Note that we use step_dummy() before step_normalize().
variable_recipe <- recipe(Grad.Rate ~ ., data = college_sub) %>%
step_nzv(all_predictors()) %>%
step_dummy(all_nominal_predictors()) %>%
step_normalize(all_numeric_predictors())STEP 3: workflow specification (model + recipe)
knn_workflow <- workflow() %>%
add_model(knn_spec) %>%
add_recipe(variable_recipe)STEP 4: estimate multiple KNN models
This code builds 50 KNN models of Grad.Rate, using 50 possible values of K ranging from 1 to 200 (roughly 25% of the sample size of 775).
It then evaluates these models with respect to their 10-fold CV MAE.
set.seed(253)
knn_models <- knn_workflow %>%
tune_grid(
grid = grid_regular(neighbors(range = c(1, 200)), levels = 50),
resamples = vfold_cv(college_sub, v = 10),
metrics = metric_set(mae)
)
knn_models %>%
autoplot()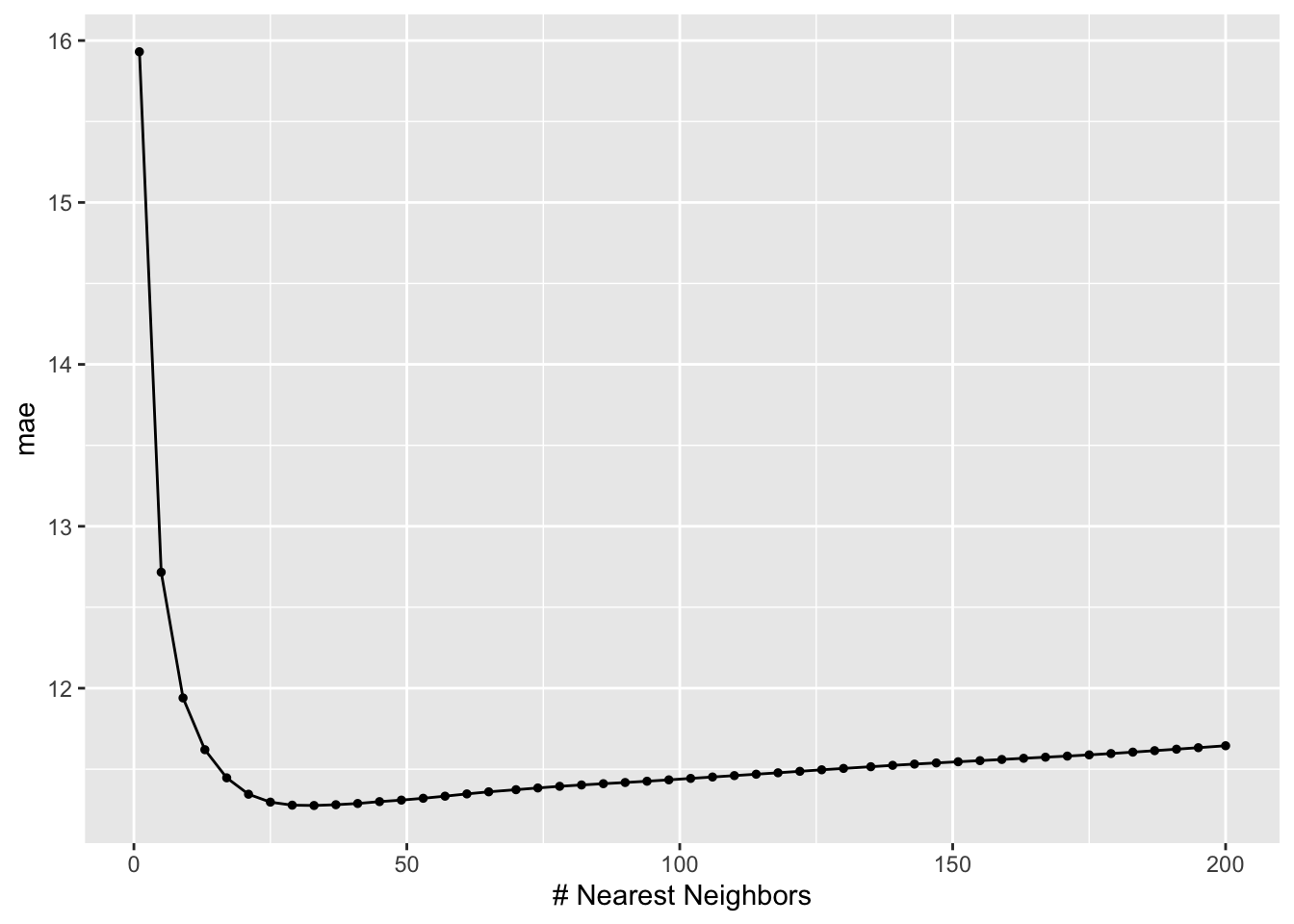
best_k <- knn_models %>%
select_best(metric = "mae")
best_k# A tibble: 1 × 2
neighbors .config
<int> <chr>
1 33 Preprocessor1_Model09# Plug in a number
knn_models %>%
collect_metrics() %>%
filter(neighbors == 33)# A tibble: 1 × 7
neighbors .metric .estimator mean n std_err .config
<int> <chr> <chr> <dbl> <int> <dbl> <chr>
1 33 mae standard 11.3 10 0.516 Preprocessor1_Model09# an equivalent but more reproducible way to do this:
knn_models %>%
collect_metrics() %>%
filter(neighbors == best_k$neighbors)# A tibble: 1 × 7
neighbors .metric .estimator mean n std_err .config
<int> <chr> <chr> <dbl> <int> <dbl> <chr>
1 33 mae standard 11.3 10 0.516 Preprocessor1_Model09final_knn <- knn_workflow %>%
finalize_workflow(parameters = best_k) %>%
fit(data = college_sub)tidy() doesn’t have estimates to give us.
# Prediction
mac_luther <- college_sub %>%
head(2)
final_knn %>%
predict(new_data = mac_luther)# A tibble: 2 × 1
.pred
<dbl>
1 78.8
2 75.7The help page tells us that select_by_one_std_err() sorts the models from most simple to most complex. For a tuning parameter p, we use:
ml_models %>% select_by_one_std_err(metric = "mae", p) (if smaller values of p indicate a simpler model)ml_models %>% select_by_one_std_err(metric = "mae", desc(p)) (if larger values of p indicate a simpler model)So for KNN we want:
simple_k <- select_by_one_std_err(knn_models, metric = "mae", desc(neighbors))
knn_models %>%
collect_metrics() %>%
filter(neighbors==simple_k$neighbors)
# To show CV results for both simple_k and best_k
knn_models %>%
collect_metrics() %>%
filter(neighbors==c(best_k$neighbors, simple_k$neighbors))Follow-up questions to consider:
select_ function would you use, and why?# a
college_sub %>%
group_by(Private) %>%
summarize(mean(Enroll))# A tibble: 2 × 2
Private `mean(Enroll)`
<fct> <dbl>
1 No 1641.
2 Yes 457.# b
ggplot(college_sub, aes(y = Grad.Rate, x = Enroll, size = Expend, color = Private)) +
geom_point(alpha = 0.5)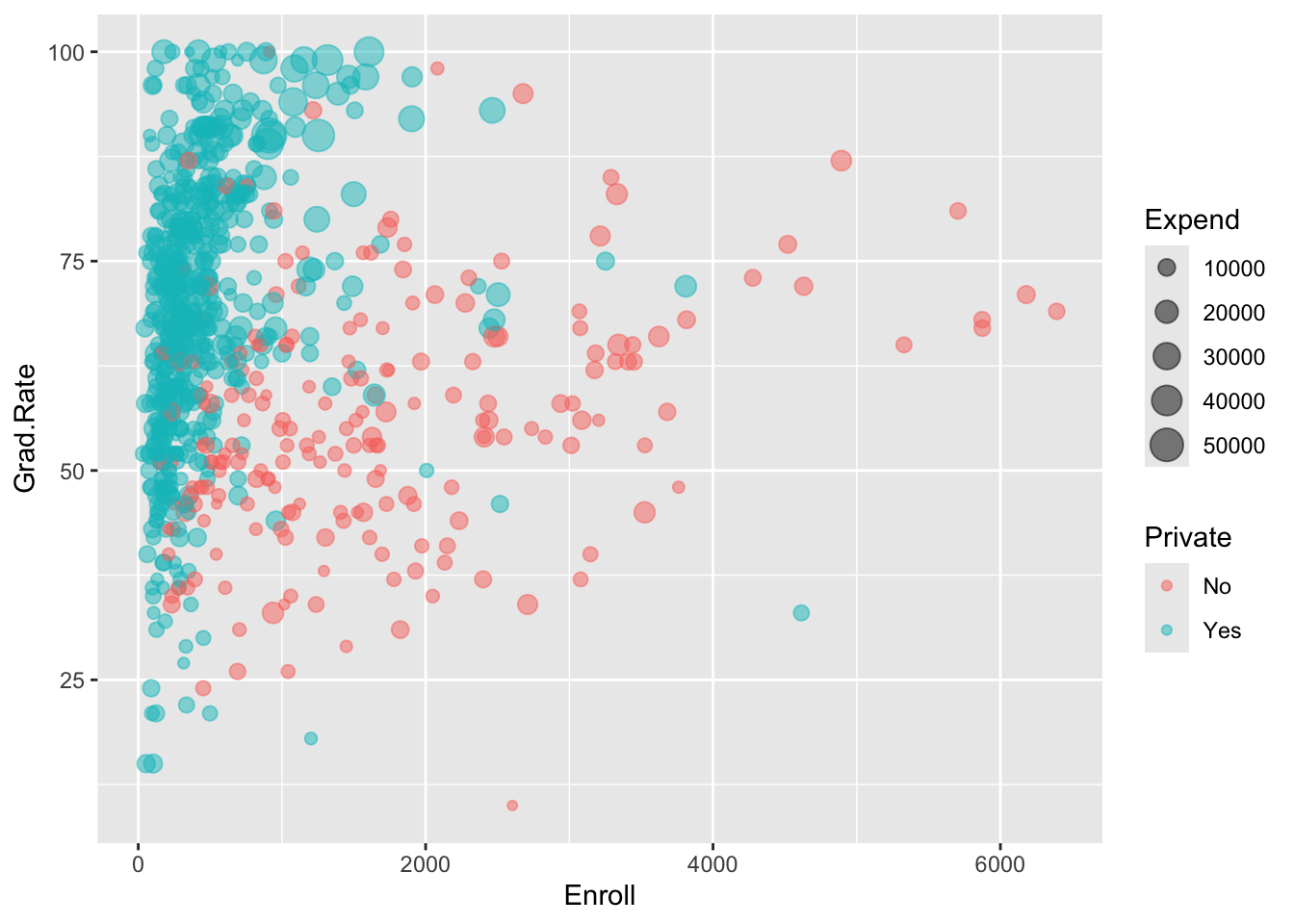
# c
college_sub %>%
filter(Enroll > 3000, Private == "Yes") Grad.Rate Expend Enroll Private
Boston University 72 16836 3810 Yes
Brigham Young University at Provo 33 7916 4615 Yes
University of Delaware 75 10650 3252 Yes